PREDOSE
PREDOSE is the acronym for PREscription Drug abuse Online Surveillance and Epidemiology, which is an inter-disciplinary project between the Ohio Center of Excellence in Knowledge-enabled Computing (Kno.e.sis) and the Center for Interventions, Treatment and Addictions Research (CITAR) at Wright State University. The overall aim of PREDOSE is to develop techniques to facilitate prescription drug abuse epidemiology, related to the illicit use of pharmaceutical opioids. PREDOSE is designed to capture the knowledge, attitudes, and behaviors of prescription drug abusers through the automatic extraction of semantic information (including entities, relationships, triples and other intelligible constructs such as sentiments, emotions, intervals, frequency, dosage, etc.) from social media. PREDOSE is the predecessor of both the eDrugTrends and iN3 projects.
Contents
[hide]News
- Researchers use social web forum data to understand nonmedical use of painkillers (article by Wright State University School of Medicine May 6, 2013)
- Semantic App Helps Researchers Understand Prescription Drug Abuse (article on Semanticweb.com June 11, 2012)
- Social Web Data on Buprenorphine Abuse Using Semantic Web Technology (article by Wright State University School of Medicine)
People
Principal Investigators: Raminta Daniulaityte, Amit P. Sheth
Co-Investigators: Robert Carlson
External Collaborators: Edward Boyer (University of Massachussetts, Amherst)
Researchers: Farahnaz Golroo, Pavan Kapanipathi, Sujan Perera, Sanjaya Wijeratne, Lu Chen, Gary A. Smith, Nishita Jaykumar, Swapnil Soni
Past Members: Delroy Cameron, Revathy Krishnamurthy, Gaurish Anand, Russel Falck (Co-Investigator), Kera Z. Watkins (Post Doc), Drashti Dave (Visiting Researcher), Pablo N. Mendes, Matthan Sink, Michael Cooney, Mandeep Singh, Pratik Desai, Mary Oberer, Kaustav Saha
Overview
The non-medical use of pharmaceutical opioids has been identified as one of the fastest growing forms of drug abuse in the U.S. The White House Office of National Drug Control Policy (ONDCP) in May 2011, launched the Epidemic: Responding to America’s Prescription Drug Abuse Crisis initiative to curb prescription drug abuse problem, mainly through education and drug monitoring programs. This White House Initiative has been prompted by recent research which associate the rise in prescription drug abuse with two important phenomena: 1) expanded pathways to heroin addiction and 2) escalating rates of accidental overdose deaths. To combat these trends, public health professionals require timely and reliable information on new and emerging patterns and trends in prescription drug abuse.
Although existing epidemiological data systems provide critically important information about drug abuse trends, they are often time-lagged. Hence, there is a critical need for content analysis platforms that could complement existing drug abuse monitoring systems and enhance the overall capacity for early identification of new and emerging patterns and trends. The World Wide Web (Web) has been identified as one of the leading data sources for detecting patterns and changes in the non-medical use of pharmaceutical and other illicit drugs. Many Web 2.0 empowered social media platforms, including web forums and tweets, provide avenues for individuals to freely share their experiences, post questions, and offer comments about various drugs. The PREDOSE project is designed to extract and analyze semantic information from online web forum discussions, as a means of detecting timely emerging patterns and trends in the non-medical use of pharmaceutical opioids. The PREDOSE project therefore has two(2) specific aims:
| Goals |
|
| Research Problem |
|
Prescription drug abuse research typically rely on manual data collection and annotation. Data are commonly gathered from interactive interviews with individual or groups of drug users. Interviews are transcribed into text, which are then manually annotated (or coded) with abstract themes. This process of qualitative coding is often facilitated using qualitative research software, such as NVivo, for Content Analysis. However, the intensive manual effort required for coding is not scalable and therefore impractical for Web-based data. Moreover, Web-based texts are fraught with grammatical errors, misspellings and slang, which can be laborious to interpret. To effectively process the large volume of abstruse heterogeneous Web-based data available from web forums, the field requires a highly automated way of extracting meaningful information from such texts, not limited to entities, sentiments, relationships and triples, |
| Approach |
|
To automate the extraction of semantic information from Web-based data, researchers from the Kno.e.sis Center at Wright State University are building information extraction techniques applied in prior research. In past research, lexical, linguistics-based, pattern-based and semantics-based processing techniques applied have been applied to automatically extract knowledge from structured biomedical texts, Wikipedia Articles, and social media (i.e., tweets). Kno.e.sis researchers have also made substantial progress in <understanding the content to: 1) identify social perceptions; 2) generate personalized information streams; 3) provide coordination and 4) identify sentiment and emotions from informal texts from MySpace, Facebook, and Twitter. Adaptations to these information processing techniques have been made to accommodate complex web forum discussions, for trend and pattern detection in prescription drug abuse research. |
| Research Plan |
|
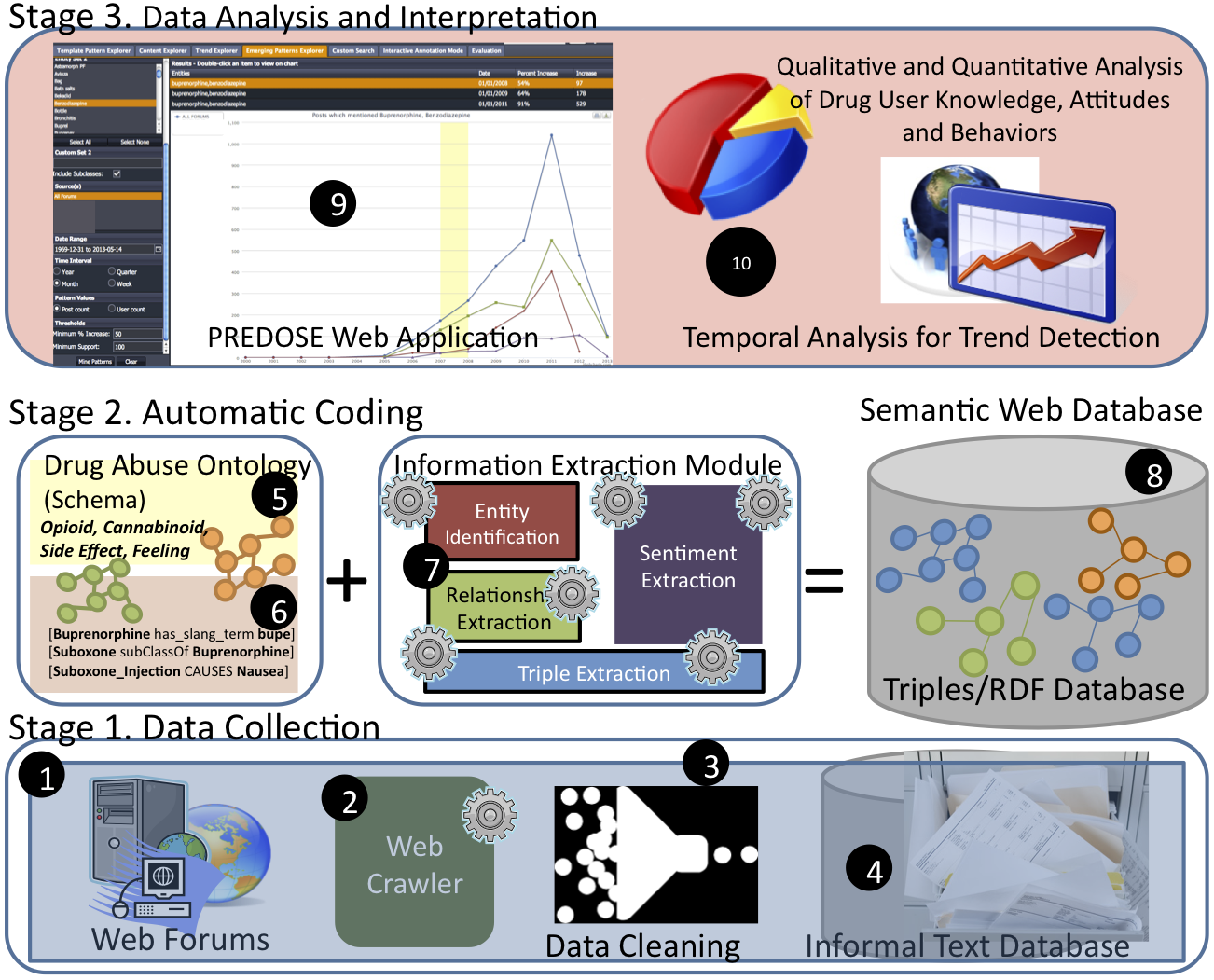
The overall research plan of the PREDOSE platform consists of three(3) stages:
|
Stage 1: Data Collection
- Web Forum Selection: The first component in the PREDOSE platform in stage 1 is for data collection. Web forums selected for the study were chosen based on the following criteria the web forum: 1) allows free discussion of psychoactive drug use; 2) contains information on illicit pharmaceutical drug use, and 3) is publicly accessible. Further, since it is important that this study collects relevant and timely information, such forums are also considered active, both in terms of number of users and diversity in topic discussions.
- Web Crawling: HTML parsers are publicly available to crawl web sites and collect data. Some of these include Nutch, Jericho HTML Parser, HTMLParser etc. In PREDOSE we use the Jericho HTML Parser to write Custom Web Crawlers to crawl data from three online web forums to obtain data for analysis.
- Data Cleaning: We sanitize the crawled HTML and decode special characters in a data cleaning phase that occurs throughout our application where necessary.
- Informal Text Database: Crawled data is stored in a MySQL database together with an index for fast retrieval. We mainly store semantic metadata in the database, based on our information extraction techniques.
Stage 2: Automatic Qualitative Coding
This is the most challenging aspect of PREDOSE. The aim is to use various information extraction techniques to extraction semantic information considered semantically equivalent to qualitative codes, from web forums. Types of extracted information include:
- Drug Abuse Ontology (DAO): We manually created a Drug Abuse Ontology (DAO) to model the prescription drug abuse domain, which is the first ontology on drug abuse in the literature. The current DAO is available online. The DAO is used to facilitate search, and it also serves as the annotation scheme for entity, relationship and triple extraction.
- Entity Identification: from web forum data is challenging because web forums discussions are informal in nature. In particular, web forum data is characterized by a proliferation of slang term references to standard drug references. We leveraged mappings for slang term to known drugs from NIDA, NDCP, Erowid, Urban Dictionary etc to enhance our domain knowledge, model. However, while such mappings are a good starting point for entity identification, the more challenging issue of entity disambiguation requires more rigorous techniques. Entity disambiguation is necessary in three scenarios: 1) standard dictionary word disambiguation (e.g. girl as Gender or the drug Cocaine); 2) word sense disambiguation (i.e., done as Methadone or the act of being done with a task) and finally 3) concept reference disambiguation (i.e. the term "Oxy" may refer to Oxycontin, Generic Oxycontin, Oxycontin OP or Oxycontin OC). We have used a combination of lexical, linguistics and semantics-based techniques to address entity identification and disambiguation: the results of which are reported in our JBI Journal article.<ref name="jbi-13"> D. Cameron, G. A. Smith, R. Daniulaityte, A. P. Sheth, D. Dave, L. Chen, G. Anand, R. Carlson, K. Z. Watkins, R. Falck. PREDOSE: A Semantic Web Platform for Drug Abuse Epidemiology using Social Media Journal of Biomedical Informatics. July 2013 ScienceDirect [PMID 23892295]</ref>
- Relationship Extraction: We have utilized a lexical and semantics-based technique for relationship identification; the details of which are reported in our JBI Journal article. <ref name ="jbi-13" />
- Triple Extraction: Previous work at Kno.e.sis have successfully implemented rule-based and probabilistic approaches to triple extraction (Ramakrishnan C, Mendes P. N. and Thomas C. Mehra P), albeit on structured biomedical literature. In another approach Thomas C and Mehra P, etc have implemented a statistical/probabilistic approach to triple extraction also on structured text. Such techniques are not likely apply to informal web forum text. Hence, we implemented a top-down pattern-based technique for triple extraction that utilizes the DAO and the declarative information extraction framework SystemT and it's implementation language AQL (Annotation Query Language), borrowing from our previous research on pattern-based information extraction from unstructured text<ref>D. Cameron, V. Bhagwan, A. P. Sheth, Towards Comprehensive Longitudinal Healthcare Data Capture. In The 1st International Workshop on the role of Semantic Web in Literature-Based Discovery, SWLBD2012 (co-located with the IEEE International Conference on Bioinformatics and Biomedicine, BIBM2012) Philadelphia PA USA, October 4, 2012, p. 241-247</ref>.
- Sentiment Extraction - We use an adaptation of the state-of-the-art sentiment extraction extraction technique developed by Chen et al<ref>Lu Chen, Wenbo Wang, Meenakshi Nagarajan, Shaojun Wang and Amit P. Sheth. Extracting Diverse Sentiment Expressions with Target-dependent Polarity from Twitter. In Proceedings of the 6th International AAAI Conference on Weblogs and Social Media (ICWSM), 2012.</ref> to extraction on-target sentiment expressions from web forum data.
- Template Pattern Identification - We use a context-free grammar <ref>D. Cameron, A. P. Sheth, N. Jaykumar, G. Anand, K.Thirunarayan, G. A. Smith. A Hybrid Approach to Finding Relevant Social Media Content for Complex Domain Specific Information Needs Journal of Web Semantics. 29: 39-52. 2014. </ref> to define the query language of strings interpretable by PREDOSE. This is a necessary task since many of the complex information needs in PREDOSE require a knowledge of ontological concepts as well as concepts not defined in ontologies such as emotion, sentiment, intensity, frequency, dosage intervals etc.
Stage 3: Data Analysis & Interpretation
In PREDOSE, we developed various components for Content Analysis. These components are included in the PREDOSE web application and the web application developed for Knowledge-Aware Search. More specifically, the PREDOSE Web Application contains components for: 1) Content Analysis and 2) spatio-temporal-thematic analysis.
- Template Pattern Explorer This is a pattern-based component for information retrieval from unstructured texts that; 1) leverages background knowledge to identify lexical variants of ontological concepts in text; 2) has the ability to semantically interpret domain specific elements (e.g. dosage, frequency of use etc) not modeled in background knowledge; 3) enables finding associations in text between template classes based on proximity, by specifying template patterns (e.g. DRUG: DOSAGE:SIDEEFFECT)
- Custom (Proximity) Search This component is a flexible lightweight extension of the Template Pattern Explorer that facilitates pattern-based search, using ontological concepts and user-specified keywords in close proximity, configurable at runtime.
- Content Explorer is a broad content exploration and annotation environment for content analysis. The exploration component enables analysis of text content restricted by 1) ontological concepts; 2) user-specified keywords; 3) specific data sources and 4) user-specified time ranges. The annotation component supports the creation of training data for information extraction tasks such as 1) entity identification and 2) sentiment extraction ubiquitous to the project.
- Trend Explorer is a component for longitudinal data analysis based on statistical aggregation of ontological concept mentions and sentiment expressions occurring text based on frequency counts and user activity.
- Emerging Patterns Explorer is an extension of the Trend Explorer for trend analysis of concomitantly occurring ontological concepts and user-specified keywords. This component is most significant because of the ability to detect spikes in discussions based on frequently co-occurring terms, unbeknownst to researchers.
A detailed description of the PREDOSE platform is available in our recently published paper in the Journal of Biomedical Informatics. <ref name="jbi-13" /> Insights into patterns and trends of Buprenorphine use are under review in the literature<ref name="cpdd-14">R. Daniulaityte, R. Carlson, D. Cameron, G. A. Smith, A. P. Sheth, When less is more: A web-based study of user beliefs about buprenorphine dosing in self-treatment of opioid withdrawal symptoms. The College on Problems of Drug Dependence CPDD 2014, San Juan, Puerto Rico, June 14-17, 2014</ref><ref name="dad-14"> R. Daniulaityte, R. Carlson, G. Brigham, D. Cameron, A. P. Sheth. "Sub is a weird drug:" A Web-based study of lay attitudes about use of buprenorphine to self-treat opioid withdrawal symptoms. American Journal of Addictions, 2015; 24(5):403-409. [PMC 4527156]</ref>
Loperamide-Withdrawal Discovery
In the early stages of the PREDOSE project we made a discovery, now reported in the literature<ref>R. Daniulaityte, R. Carlson, R. Falck, D. Cameron, S. Perera, L. Chen, A. P. Sheth. "I Just Wanted to Tell You That Loperamide WILL WORK": A Web-Based Study of Extra-Medical Use of Loperamide. Journal of Drug and Alcohol Dependence. 130(1-3): 241-244, 2013. ScienceDirect, [PMID 23201175]</ref> <ref>R. Daniulaityte, R. Carlson, R. Falck, D. Cameron, S. Perera, L. Chen, A. P. Sheth. A Web-Based Study of Self-Treatment of Opioid Withdrawal Symptoms with Loperamide. The College on Problems of Drug Dependence CPDD 2012, Palm Springs, CA USA, June 9-14, 2012.</ref>.
Based on the lexical and semantics-based techniques for entity identification various datasets were isolated according to drug mentions, based on mapping slang references to standard concepts. In one dataset, it was observed that users reported taking the anti-diarrhea treatment drug Loperamide (sold over the counter in Imodium) to self-medicate from withdrawal symptoms. The opioid addictions treatment drugs Buprenorphine and Methadone are commonly prescribed for treatment of withdrawal symptoms. Until now, it was unknown that Loperamide, can be (and is being) used for the same purpose. Which is more, it was observed that users reported the possibility of mild psychoactive (opiated) effects from megadosing - which is the practice of taking severely excessive amounts of a drug.
PREDOSE Live
http://knoesis-hpco.cs.wright.edu/predose/ [Video Demo]
http://knoesis-hpco.cs.wright.edu/knowledge-aware-search [Video Demo]
Publications
- R. Daniulaityte, R. Carlson, G. Brigham, D. Cameron, A. P. Sheth. A Study of Social Web Data on Buprenorphine Abuse Using Semantic Web Technology. Final Report.
- D. Cameron, G. A. Smith, R. Daniulaityte, A. P. Sheth, D. Dave, L. Chen, G. Anand, R. Carlson, K. Z. Watkins, R. Falck. PREDOSE: A Semantic Web Platform for Drug Abuse Epidemiology using Social Media Journal of Biomedical Informatics. July 2013
Related
- Researchers use social web forum data to understand nonmedical use of painkillers
- U.S. Targeting Prescription Drug Abuse
- Twitter Helps Determine "Morning People" and "Night Owls"
- Knowledge-Aware-Search
Related Projects
- Innovative NIDA National Early Warning Sysetm Network (iN3)
- EDrugTrends
- Hazards SEES: Social and Physical Sensing Enabled Decision Support
- DAO
Funding
This project was initially sponsored by the National Institutes of Health (NIH) Grant No. R21 DA030571-01A1 awarded to the Ohio Center of Excellence in Knowledge-enabled Computing (Kno.e.sis) and the Center for Interventions, Treatment and Addictions Research (CITAR) titled: A Study of Social Web Data on Buprenorphine Abuse using Semantic Web Technology. It is continued under National Institutes on Drug Abuse (NIDA) Grant No. R56DA038366-01, titled: NIDA National Early Warning System Network (iN3): An Innovative Approach (wiki page). Any opinions, findings, conclusions or recommendations expressed in this material are those of the investigator(s) and do not necessarily reflect the views of the National Institutes of Health.
Contact: Farahnaz Golroo