Parasite Experiment ontology
The ontology comprehensively models the processes, instruments, parameters, and sample details that will be used to annotate experimental results with “provenance” metadata (derivation history of results). For example, given a drug selected sample (a sample that has been grown under selective drug pressure. source: Parasite Experiment ontology), details about the drug, drug concentration, the duration for the drug selection process, and the transfected sample used as input to the drug selection process are modeled in the ontology. This extensive provenance metadata modeled in this ontology will enable publication of results in journals, conferences with details of the method used to arrive at the result. Parasite Experiment ontology currently has 71 classes and 15 properties with a DL expressivity of ALCHF(D) and incorporates concepts defined in Parasite Life Cycle ontology to maximize re-use of existing ontological resources.
|
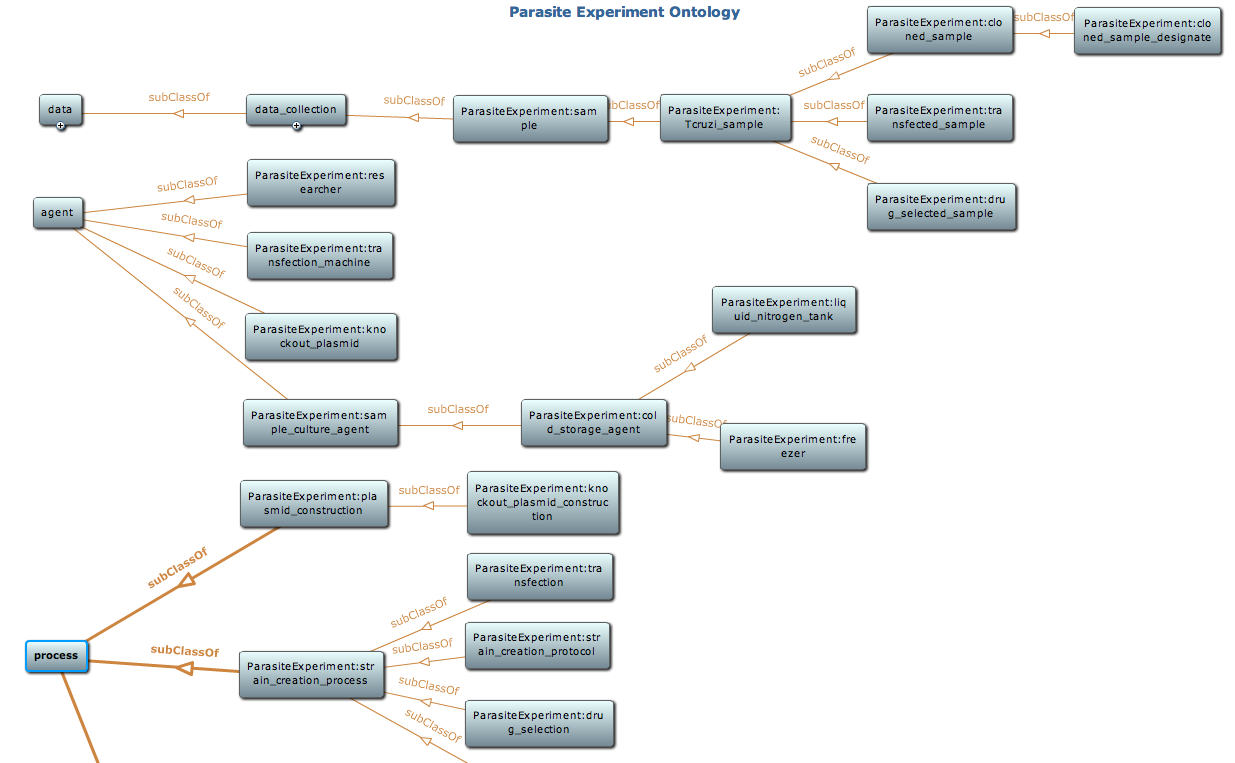 Snapshot of class hierarchy of the parasite experiment ontology using Protege toolkit. Detail of this ontology is available at NCBO BioPortal |
List of ontology classes
Collaborating Institutions
We invite new members to collaborate with us in extending this ontology.
Feedback/Comments
Please add feedback/comments in Discussion Section on the top of this wiki page.