Difference between revisions of "Parasite Experiment ontology"
| Line 14: | Line 14: | ||
{{#tree:id=OwlTree|openlevels=1|root=Owl:Thing| | {{#tree:id=OwlTree|openlevels=1|root=Owl:Thing| | ||
*agent | *agent | ||
| + | **[[Data_Source|data_source]] | ||
**[[Freezer|freezer]] | **[[Freezer|freezer]] | ||
**[[Knockout_Plasmid|knockout_plasmid]] | **[[Knockout_Plasmid|knockout_plasmid]] | ||
| − | |||
**[[Microarray_Plate_Reader|microarray_plate_reader]] | **[[Microarray_Plate_Reader|microarray_plate_reader]] | ||
| + | **[[Researcher|researcher]] | ||
**[[Transfection_Machine|transfection_machine]] | **[[Transfection_Machine|transfection_machine]] | ||
*data | *data | ||
**data_collection | **data_collection | ||
| + | ***[[GO_Term|go_term]] | ||
***[[Peptide_count|peptide_count]] | ***[[Peptide_count|peptide_count]] | ||
***[[Phenotype|phenotype]] | ***[[Phenotype|phenotype]] | ||
***[[Protein_group_number|protein_group_number]] | ***[[Protein_group_number|protein_group_number]] | ||
| − | ***[[sample]] | + | ***[[SAM_result|sam_result]] |
| + | ***[[Sample|sample]] | ||
****[[Tcruzi_Lifecyclestage_Subsample|Tcruzi_lifecyclestage_subsample]] | ****[[Tcruzi_Lifecyclestage_Subsample|Tcruzi_lifecyclestage_subsample]] | ||
****[[Tcruzi_Sample|Tcruzi_sample]] | ****[[Tcruzi_Sample|Tcruzi_sample]] | ||
| Line 49: | Line 52: | ||
****[[Culture_Parameter|culture_parameter]] | ****[[Culture_Parameter|culture_parameter]] | ||
*****[[Temperature_Condition|temperature_condition]] | *****[[Temperature_Condition|temperature_condition]] | ||
| − | ****[[drug]] | + | ****[[Drug|drug]] |
****[[Drug_Attribute|drug_attribute]] | ****[[Drug_Attribute|drug_attribute]] | ||
*****[[Drug_Concentration|drug_concentration]] | *****[[Drug_Concentration|drug_concentration]] | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
****[[Nucleic_Acid|nucleic_acid]] | ****[[Nucleic_Acid|nucleic_acid]] | ||
| + | ****[[Primer|primer]] | ||
| + | *****[[2_Prime_Primer|2_prime_primer]] | ||
| + | ******[[2_Prime_Forward_Primer|2_prime_forward_primer]] | ||
| + | ******[[2_Prime_Reverse_Primer|2_prime_reverse_primer]] | ||
| + | *****[[3_Prime_Primer|3_prime_primer]] | ||
| + | ******[[3_Prime_Forward_Primer|3_prime_forward_primer]] | ||
| + | ******[[3_Prime_Reverse_Primer|3_prime_reverse_primer]] | ||
****[[Purity|purity]] | ****[[Purity|purity]] | ||
****[[Transfection_Buffer|transfection_buffer]] | ****[[Transfection_Buffer|transfection_buffer]] | ||
| − | ****[[ | + | ****[[Trg_Intra_Lab_Parameters|trg_intra_lab_parameters]] |
| − | ****[[ | + | ****[[Host_Organism|host_organism]] |
| − | ****[[ | + | ****[[Lifecycle_Stage|lifecycle_stage]] |
| + | *****[[Amastigote|amastigote]] | ||
| + | ******[[Leishmania_SP_Amastigote|leishmania_sp_amastigote]] | ||
| + | ******[[Trypanosoma_Cruzi_Amastigote|trypanosoma_cruzi_amastigote]] | ||
| + | *****[[Epimastigote|epimastigote]] | ||
| + | ******[[Trypanosoma_Brucei_Epimastigote|trypanosoma_cruzi_epimastigote]] | ||
| + | *****[[Promastigote|promastigote]] | ||
| + | ******[[Metacyclic_Promastigote|metacyclic_promastigote]] | ||
| + | ******[[Procyclic_Promastigote|procyclic_promastigote]] | ||
| + | *****[[Trypomastigote|trypomastigote]] | ||
| + | ******[[Blood_Stream_Form_Trypomastigote|blood_stream_form_trypomastigote]] | ||
| + | ******[[Metacyclic_Trypomastigote|metacyclic|trypomastigote]] | ||
| + | *******[[Trypanosoma_brucei_Metacyclic_Trypomastigote|trypanosoma_brucei_metacyclic_trypomastigote]] | ||
| + | *******[[Trypanosoma_Cruzi_Metacyclic_Trypomastigote|trypanosoma_cruzi_metacyclic_trypomastigote]] | ||
| + | ******[[Procyclic_Trypomastigote|procyclic_trypomastigote]] | ||
| + | ******[[Trypanosoma_Brucei_Trypomastigote|trypanosoma_brucei_trypomastigote]] | ||
| + | ******[[Trypanosoma_Cruzi_Trypomastigote|trypanosoma_cruzi_trypomastigote]] | ||
****[[organism]] | ****[[organism]] | ||
****[[Organism_Strain|organism_strain]] | ****[[Organism_Strain|organism_strain]] | ||
| Line 73: | Line 89: | ||
****[[Genome|genome]] | ****[[Genome|genome]] | ||
*****[[Transfected_Genome|transfected_genome]] | *****[[Transfected_Genome|transfected_genome]] | ||
| − | + | ****[[Region|region]] | |
| − | ******[[ | + | *****[[Gene|gene]] |
| − | ******[[ | + | *****[[Intergenic_Region|intergenic_region]] |
| − | ******[[ | + | ******[[2_Prime_Intergenic_Region|2_prime_intergenic_region]] |
| + | *******[[2_Prime_Intergenic_Forward_Region|2_prime_intergenic_forward_region]] | ||
| + | *******[[2_Prime_Intergenic_Reverse_Region|2_prime_intergenic_reverse_region]] | ||
| + | ******[[3_Prime_Intergenic_Region|3_prime_intergenic_region]] | ||
| + | *******[[3_Prime_Intergenic_Forward_Region|3_prime_intergenic_forward_region]] | ||
| + | *******[[3_Prime_Intergenic_Reverse_Region|3_prime_intergenic_reverse_region]] | ||
| + | ****[[Gene_Function|gene_function]] | ||
***spatial_parameter | ***spatial_parameter | ||
****[[Freezer_Location|freezer_location]] | ****[[Freezer_Location|freezer_location]] | ||
| Line 82: | Line 104: | ||
***temporal_parameter | ***temporal_parameter | ||
****[[Date_Time_Description|DateTimeDescription]] | ****[[Date_Time_Description|DateTimeDescription]] | ||
| + | *****[[DateCompleted|dateCompleted]] | ||
| + | *****[[DateOfLastModification|dateOfLastModification]] | ||
| + | *****[[DateStarted|dateStarted]] | ||
*process | *process | ||
**[[Gene_Knockout_Process|gene_knockout_process]] | **[[Gene_Knockout_Process|gene_knockout_process]] | ||
| Line 122: | Line 147: | ||
<tr valign="top"> | <tr valign="top"> | ||
<td> | <td> | ||
| + | <ul> | ||
| + | <li>[[2 Prime Forward Primer]]</li> | ||
| + | <li>[[2 Prime Intergenic Forward Region]]</li> | ||
| + | <li>[[2 Prime Intergenic Region]]</li> | ||
| + | <li>[[2 Prime Intergenic Reverse Region]]</li> | ||
| + | <li>[[2 Prime Primer]]</li> | ||
| + | <li>[[2 Prime Reverse Primer]]</li> | ||
| + | <li>[[3 Prime Forward Primer]]</li> | ||
| + | <li>[[3 Prime Intergenic Forward Region]]</li> | ||
| + | <li>[[3 Prime Intergenic Region]]</li> | ||
| + | <li>[[3 Prime Intergenic Reverse Region]]</li> | ||
| + | <li>[[3 Prime Primer]]</li> | ||
| + | <li>[[3 Prime Reverse Primer]]</li> | ||
| + | </ul> | ||
;A | ;A | ||
<ul> | <ul> | ||
| − | + | <li> [[Accession Number]]</li> | |
| − | + | ||
<li> [[Amastigote]]</li> | <li> [[Amastigote]]</li> | ||
</ul> | </ul> | ||
| + | |||
| + | ;B | ||
| + | <ul> | ||
| + | <li>[[Blood Stream Form Trypomastigote]]</li> | ||
| + | </ul> | ||
;C | ;C | ||
<ul> | <ul> | ||
| − | + | <li>[[Cell Cloning]]</li> | |
<li>[[Chromosome]]</li> | <li>[[Chromosome]]</li> | ||
| − | + | <li>[[Cloned Sample]]</li> | |
| − | + | <li>[[Cloned Sample Designate]]</li> | |
<li>[[Culture Medium]]</li> | <li>[[Culture Medium]]</li> | ||
<li>[[Culture Parameter]]</li> | <li>[[Culture Parameter]]</li> | ||
| Line 141: | Line 184: | ||
;D | ;D | ||
<ul> | <ul> | ||
| + | <li>[[Data Source]]</li> | ||
| + | <li>[[Date Completed]]</li> | ||
| + | <li>[[Date of Last Modification]]</li> | ||
| + | <li>[[Date Started]]</li> | ||
<li>[[Date Time Description]]</li> | <li>[[Date Time Description]]</li> | ||
<li>[[Drug]]</li> | <li>[[Drug]]</li> | ||
<li>[[Drug Attribute]]</li> | <li>[[Drug Attribute]]</li> | ||
| − | + | <li>[[Drug Concentration]]</li> | |
| − | + | <li>[[Drug Resistant Plasmid]]</li> | |
| − | + | <li>[[Drug Selected Sample]]</li> | |
| − | + | <li>[[Drug Selection]]</li> | |
</ul> | </ul> | ||
;E | ;E | ||
<ul> | <ul> | ||
<li>[[Epimastigote]]</li> | <li>[[Epimastigote]]</li> | ||
| − | |||
| − | |||
</ul> | </ul> | ||
;F | ;F | ||
<ul> | <ul> | ||
| − | |||
<li>[[Freezer]]</li> | <li>[[Freezer]]</li> | ||
<li>[[Freezer Location]]</li> | <li>[[Freezer Location]]</li> | ||
| Line 168: | Line 212: | ||
<ul> | <ul> | ||
<li>[[Gene]]</li> | <li>[[Gene]]</li> | ||
| − | <li>[[Gene Function]]</li> | + | <li>[[Gene Function]]</li> |
| − | <li>[[Gene Knockout | + | <li>[[Gene Knockout Process]]</li> |
<li>[[Genome]]</li> | <li>[[Genome]]</li> | ||
| + | <li>[[GO Term]]</li> | ||
</ul> | </ul> | ||
;H | ;H | ||
<ul> | <ul> | ||
| − | + | <li>[[Host organism]]</li> | |
| − | + | <li>[[Hybridisation]]</li> | |
</ul> | </ul> | ||
;I | ;I | ||
<ul> | <ul> | ||
| − | + | <li>[[Intergenic Region]]</li> | |
</ul> | </ul> | ||
| Line 187: | Line 232: | ||
<li>[[Kinetoplast Position]]</li> | <li>[[Kinetoplast Position]]</li> | ||
<li>[[Knockout Construct Plasmid]]</li> | <li>[[Knockout Construct Plasmid]]</li> | ||
| − | + | <li>[[Knockout Plasmid]]</li> | |
<li>[[Knockout Plasmid Construction]]</li> | <li>[[Knockout Plasmid Construction]]</li> | ||
<li>[[Knockout Project Protocol]]</li> | <li>[[Knockout Project Protocol]]</li> | ||
| Line 195: | Line 240: | ||
;L | ;L | ||
<ul> | <ul> | ||
| − | + | <li>[[Labelling]]</li> | |
<li>[[Laboratory Experiment Parameter]]</li> | <li>[[Laboratory Experiment Parameter]]</li> | ||
| − | + | <li>[[Leishmania SP Amastigote]]</li> | |
| − | <li>[[Log base2 Ratio]]</li> </ul> | + | <li>[[Lifecycle stage]]</li> |
| + | <li>[[Log base2 Ratio]]</li> | ||
| + | </ul> | ||
</td> | </td> | ||
<td> | <td> | ||
;M | ;M | ||
<ul> | <ul> | ||
| − | + | <li>[[Metacyclic_Promastigote]]</li> | |
| − | + | <li>[[Metacyclic_Trypomastigote]]</li> | |
| − | <li>[[Microarray | + | <li>[[Microarray Analysis]]</li> |
<li>[[Microarray Oligonucleotide]]</li> | <li>[[Microarray Oligonucleotide]]</li> | ||
| − | + | <li>[[Microarray Plate Reader]]</li> | |
</ul> | </ul> | ||
;N | ;N | ||
<ul> | <ul> | ||
| − | + | <li>[[Nucleic Acid]]</li> | |
</ul> | </ul> | ||
| Line 224: | Line 271: | ||
;P | ;P | ||
<ul> | <ul> | ||
| − | + | <li>[[Peptide count]]</li> | |
| − | + | <li>[[Phenotype]]</li> | |
| − | + | <li>[[Plasmid]]</li> | |
| − | + | <li>[[Plasmid Construction]]</li> | |
| − | <li>[[ | + | <li>[[Polymerase Chain Reaction]]</li> |
| − | + | <li>[[Prehybridisation]]</li> | |
| − | + | <li>[[Primer]]</li> | |
| − | + | <li>[[Priority]]</li> | |
| − | <li>[[Probability]]</li> | + | <li>[[Probability]]</li> |
| + | <li>[[Procyclic Promastigote]]</li> | ||
| + | <li>[[Procyclic Trypomastigote]]</li> | ||
<li>[[Promastigote]]</li> | <li>[[Promastigote]]</li> | ||
<li>[[Protein group number]]</li> | <li>[[Protein group number]]</li> | ||
| Line 246: | Line 295: | ||
<ul> | <ul> | ||
<li>[[Region]]</li> | <li>[[Region]]</li> | ||
| − | <li>[[Researcher | + | <li>[[Researcher]]</li> |
| − | + | ||
</ul> | </ul> | ||
;S | ;S | ||
<ul> | <ul> | ||
| − | + | <li>[[SAM Result]]</li> | |
| + | <li>[[Sample]]</li> | ||
<li>[[Sample Characterization]]</li> | <li>[[Sample Characterization]]</li> | ||
<li>[[Scanning]]</li> | <li>[[Scanning]]</li> | ||
<li>[[Sequence Coverage]]</li> | <li>[[Sequence Coverage]]</li> | ||
| − | + | <li>[[Sequence Extraction]]</li> | |
| − | + | <li>[[Signal Intensity]]</li> | |
<li>[[Spectral Count]]</li> | <li>[[Spectral Count]]</li> | ||
<li>[[Stage Purity Determination]]</li> | <li>[[Stage Purity Determination]]</li> | ||
| Line 262: | Line 311: | ||
<li>[[Statistical Measure]]</li> | <li>[[Statistical Measure]]</li> | ||
<li>[[Status]]</li> | <li>[[Status]]</li> | ||
| − | |||
<li>[[Strain Creation Process]]</li> | <li>[[Strain Creation Process]]</li> | ||
<li>[[Strain Creation Protocol]]</li> | <li>[[Strain Creation Protocol]]</li> | ||
| Line 270: | Line 318: | ||
<li>[[Target Region Plasmid]]</li> | <li>[[Target Region Plasmid]]</li> | ||
<li>[[Tcruzi Lifecyclestage Subsample]]</li> | <li>[[Tcruzi Lifecyclestage Subsample]]</li> | ||
| − | + | <li>[[Tcruzi Sample]]</li> | |
| − | + | ||
<li>[[Temperature Condition]]</li> | <li>[[Temperature Condition]]</li> | ||
<li>[[Transcriptome Process]]</li> | <li>[[Transcriptome Process]]</li> | ||
| − | |||
<li>[[Transfected Genome]]</li> | <li>[[Transfected Genome]]</li> | ||
<li>[[Transfected Sample]]</li> | <li>[[Transfected Sample]]</li> | ||
| − | + | <li>[[Transfection]]</li> | |
| − | + | <li>[[Transfection Buffer]]</li> | |
| − | + | <li>[[Transfection Machine]]</li> | |
| + | <li>[[TRG Intra Lab Parameters]]</li> | ||
| + | <li>[[Trypanosoma Brucei Epimastigote]]</li> | ||
| + | <li>[[Trypanosoma Brucei Metacyclic Trypomastigote]]</li> | ||
| + | <li>[[Trypanosoma Brucei Trypomastigote]]</li> | ||
| + | <li>[[Trypanosoma Cruzi Amastigote]]</li> | ||
| + | <li>[[Trypanosoma Cruzi Metacyclic Trypomastigote]]</li> | ||
| + | <li>[[Trypanosoma Cruzi Trypomastigote]]</li> | ||
<li>[[Trypomastigote]]</li> | <li>[[Trypomastigote]]</li> | ||
</ul> | </ul> | ||
Revision as of 16:50, 11 January 2011
| What is Parasite Experiment Ontology (PEO)? |
|
The ontology comprehensively models the processes, instruments, parameters, and sample details that will be used to annotate experimental results with “provenance” metadata (derivation history of results). For example, given a drug selected sample (a sample that has been grown under selective drug pressure. source: Parasite Experiment ontology), details about the drug, drug concentration, the duration for the drug selection process, and the transfected sample used as input to the drug selection process are modeled in the ontology. This extensive provenance metadata modeled in this ontology will enable publication of results in journals, conferences with details of the method used to arrive at the result. PEO has 110 classes and 27 properties with a logic expressivity of ALCHQ(D) and incorporates concepts defined in Parasite Life Cycle ontology (PLO) to maximize reuse of existing ontological resources. The new version of PEO (ver 0.2) was released for listing and public use at NCBO’s BioPortal in December 2009. |
|
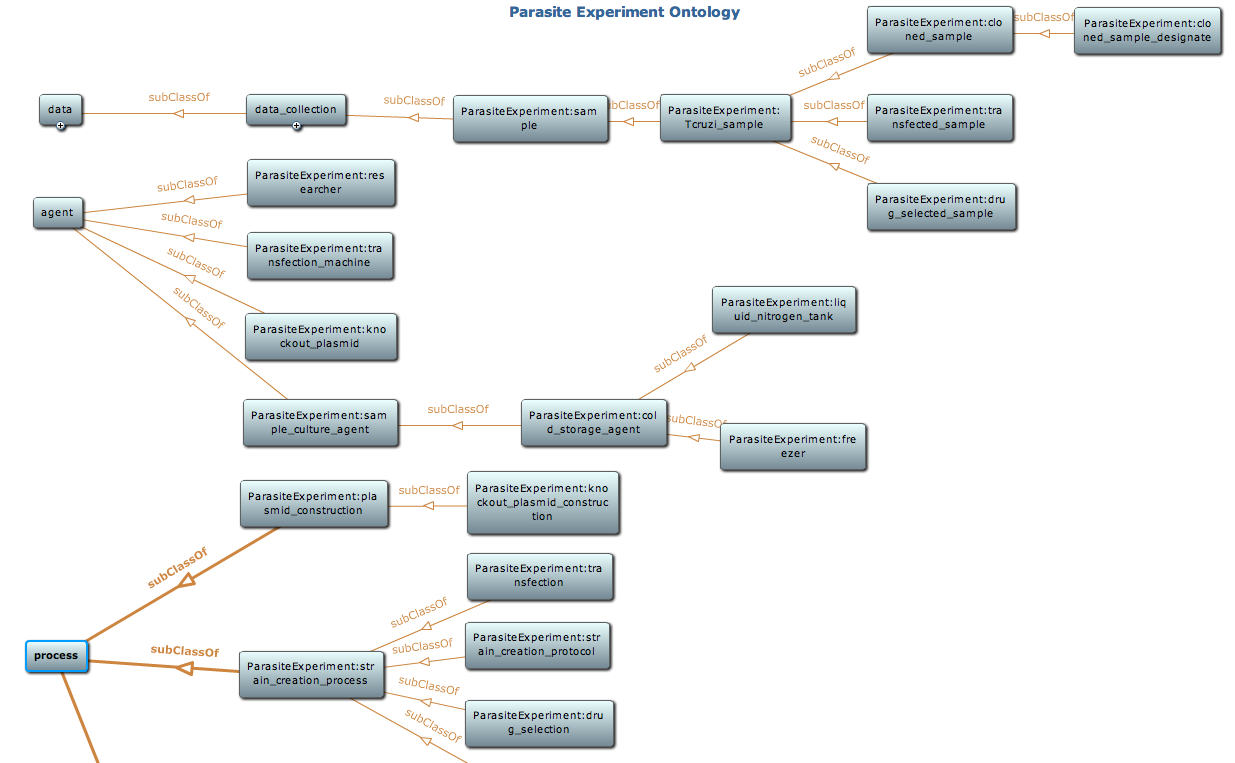 Snapshot of class hierarchy of the parasite experiment ontology using Protege toolkit. Detail of this ontology is available at NCBO BioPortal |
| Feedback/Comments |
|
Please add feedback/comments in Discussion Section on the top of this wiki page. |