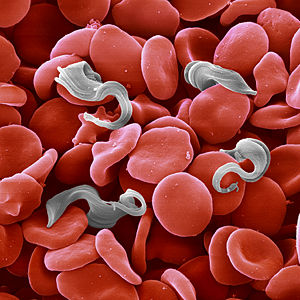
 Parasitic Trypanosoma brucei surrounded by red blood cells in a smear of infected blood. (Copyright: Jürgen Berger and Dr. Peter Overath, Max Planck Institute for Developmental Biology, Tübingen.) Trypanosoma cruzi is a protozoan parasite and a relative of other human pathogens that cause African sleeping sickness and leishmaniasis. Approximately 18 million people in Latin America are infected with this parasite, and as many as 40% of these are predicted to eventually suffer from Chagas disease, which is the leading cause of heart disease and sudden death in middle-aged adults in the region. Currently, there is a single manufactured drug for treatment of the infection; but this drug has severe side effects and is also carcinogenic. Diagnosis of the infection is poor and there are no vaccines for prevention of the infection. Moreover, T. cruzi presents a level of complexity that, though not unique, is substantial.
Research in many biological systems, particularly those involving infectious agents, continues to experience an enormous increase in information with the sequencing of genomes, the use of expression profiling, and the completion of proteomic analyses. For example, annotated genomes and comparative analyses of two T. cruzi related kinetoplastids, Trypanosoma brucei and Leishmania major, and a whole organism (all life-cycle stages) proteome analysis of T. cruzi have been published. Data from other sequencing projects in T. cruzi, including ESTs, preexisting and ongoing expression profiling (DNA microarray) and proteome analyses are also available. These dynamic datasets, addressing different but logically related aspects, are distributed over multiple databases that undergo frequent additions and curation. It is difficult to integrate these datasets manually from multiple sources. Therefore, we propose to use ontology-driven technology to integrate local and external datasets to answer biological questions at mulitple level of granularity. Project Page
|
