Difference between revisions of "Provenir Ontology"
Satyasahoo (Talk | contribs) m |
|||
| (14 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| − | <!-- | + | <!--begin table --> |
| + | {| style="width:100%; vertical-align:top;" | ||
| + | <!--begin left panel --> | ||
| + | | style="width:60%; border:1px solid #cef2e0; background:#cef2e0; vertical-align:top; color:#000;" | | ||
| + | {| style="width:100%; vertical-align:top;" | ||
| + | |||
| + | <!--start section left 1 --> | ||
| + | |- | ||
| + | | | ||
{{block | {{block | ||
| − | |title= What is Provenir Ontology (PO)? | + | |title= <span style="color:#ffffff">What is Provenir Ontology (PO)?</span> |
| − | |title_background=# | + | |title_background=#008b8b |
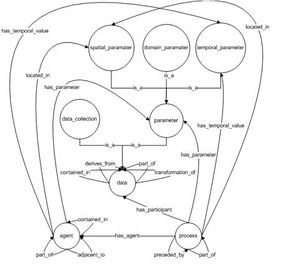
| − | |content= A reference ontology for modeling domain-specific provenance[[Image:The_provenir_ontology_schema.jpg | right | | + | |content= A reference ontology for modeling domain-specific provenance[[Image:The_provenir_ontology_schema.jpg | right | thumb | Provenir Ontology Schema]] |
Provenance, from the French word ‘provenir’ meaning to come from, describes the lineage of an entity. Provenance is critical information in eScience to accurately interpret scientific results. Information provenance has been recognized as a hard problem in computing science (British Computing Society, 2004), and many research issues in provenance are yet to be addressed. For example, a common provenance model to facilitate interoperability of provenance metadata and to support analysis using inferencing rules has not been defined. | Provenance, from the French word ‘provenir’ meaning to come from, describes the lineage of an entity. Provenance is critical information in eScience to accurately interpret scientific results. Information provenance has been recognized as a hard problem in computing science (British Computing Society, 2004), and many research issues in provenance are yet to be addressed. For example, a common provenance model to facilitate interoperability of provenance metadata and to support analysis using inferencing rules has not been defined. | ||
We introduce the provenir ontology as a common provenance model, which forms the core component of a modular approach to provenance management framework in eScience. Domain-specific details are an important component of provenance representation. But, a single monolithic provenance ontology that models all possible details from different domains (biology, marine sciences, and astronomy) is clearly not feasible. Hence, our proposed modular approach involves integrated use of multiple ontologies, each modeling provenance metadata specific to a particular domain (for example, the ProPreO ontology represents proteomics domain-specific provenance[1]). These multiple ontologies will use the provenir ontology as the common reference model, hence making it easier for their associated instances to be interoperable. This modular framework represents a scalable and flexible approach to provenance modeling that can be adapted to the specific requirement of different domains. In the next two Sections, we describe the classes and the properties in the provenir ontology (Figure 1). | We introduce the provenir ontology as a common provenance model, which forms the core component of a modular approach to provenance management framework in eScience. Domain-specific details are an important component of provenance representation. But, a single monolithic provenance ontology that models all possible details from different domains (biology, marine sciences, and astronomy) is clearly not feasible. Hence, our proposed modular approach involves integrated use of multiple ontologies, each modeling provenance metadata specific to a particular domain (for example, the ProPreO ontology represents proteomics domain-specific provenance[1]). These multiple ontologies will use the provenir ontology as the common reference model, hence making it easier for their associated instances to be interoperable. This modular framework represents a scalable and flexible approach to provenance modeling that can be adapted to the specific requirement of different domains. In the next two Sections, we describe the classes and the properties in the provenir ontology (Figure 1). | ||
}} | }} | ||
| − | <!--end section 1 | + | <!-- end section left 1 --> |
| − | + | ||
| − | + | ||
| + | <!--start section left 2 --> | ||
| + | |- | ||
| + | | | ||
{{block | {{block | ||
| − | |title= Classes of Provenir Ontology (PO) | + | |title= <span style="color:#ffffff">Classes of Provenir Ontology (PO)</span> |
| − | |title_background=# | + | |title_background=#008b8b |
|content= To represent provenance metadata classes we use the two well defined, primitive concepts of “occurrent” and “continuant” from philosophical ontology[2]. Continuant is defined as “… entities which endure, or continue to exist, through time while undergoing different sorts of changes, including changes of place” [2]. Occurrent is defined as “…entities that unfold themselves in successive temporal phases”[2]. | |content= To represent provenance metadata classes we use the two well defined, primitive concepts of “occurrent” and “continuant” from philosophical ontology[2]. Continuant is defined as “… entities which endure, or continue to exist, through time while undergoing different sorts of changes, including changes of place” [2]. Occurrent is defined as “…entities that unfold themselves in successive temporal phases”[2]. | ||
We define three base classes in the provenir ontology representing the primary components of provenance, that is, “data”, “agent” and “process”. The two base classes, “data” and “agents” are defined as specialization (sub-class) of continuant class. The third base class “process” is a synonym of occurrent. We present the definition of each class capturing inheritance relationship: | We define three base classes in the provenir ontology representing the primary components of provenance, that is, “data”, “agent” and “process”. The two base classes, “data” and “agents” are defined as specialization (sub-class) of continuant class. The third base class “process” is a synonym of occurrent. We present the definition of each class capturing inheritance relationship: | ||
| Line 29: | Line 38: | ||
:::* domain_parameter: The domain_parameter class is used to model domain-specific parameters (for example, tolerable salinity levels for ocean buoys). | :::* domain_parameter: The domain_parameter class is used to model domain-specific parameters (for example, tolerable salinity levels for ocean buoys). | ||
}} | }} | ||
| − | <!--end section 2 --> | + | <!-- end section left 2 --> |
| − | <!--start section 3 --> | + | <!--start section left 3 --> |
| − | + | |- | |
| + | | | ||
{{block | {{block | ||
| − | |title= Properties of Provenir Ontology (PO) | + | |title= <span style="color:#ffffff">Properties of Provenir Ontology (PO)</span> |
| − | |title_background=# | + | |title_background=#008b8b |
|content= In this section, we define a set of foundational properties in the provenir ontology. | |content= In this section, we define a set of foundational properties in the provenir ontology. | ||
Instead of defining a new set of properties, we adapt the properties defined in the Relation ontology (RO) from the Open Biomedical Ontologies (OBO) Foundry: | Instead of defining a new set of properties, we adapt the properties defined in the Relation ontology (RO) from the Open Biomedical Ontologies (OBO) Foundry: | ||
| Line 50: | Line 60: | ||
:::* located_in – An instance of data or agent is associated with exactly one spatial region that is its exact location at given instance of time. In provenir ontology, this relation has two domain class agent and data_collection with spatial_parameter as range class. | :::* located_in – An instance of data or agent is associated with exactly one spatial region that is its exact location at given instance of time. In provenir ontology, this relation has two domain class agent and data_collection with spatial_parameter as range class. | ||
}} | }} | ||
| − | <!--end section 3 --> | + | <!-- end section left 3 --> |
| − | + | |}<!-- end left panel --> | |
| − | [http://knoesis.wright.edu/library/ontologies/provenir/provenir.owl Provenir ontology schema (OWL file)] | + | <!-- begin right panel --> |
| + | | style="width:40%; border:1px solid #ffffcc; background:#f5faff; vertical-align:top;" | <!--right panel --> | ||
| + | {| style="width:100%; vertical-align:top;" | ||
| + | |||
| + | <!--start section right 1 --> | ||
| + | |- | ||
| + | | | ||
| + | {{block | ||
| + | |title= <span style="color:#ffffff">Quick Links</span> | ||
| + | |title_background=#556b2f | ||
| + | |content= | ||
| + | * Applications of Provenir Ontology in different domains | ||
| + | **[[Biomedical Sciences]] | ||
| + | **[[Oceanography]] | ||
| + | **[http://knoesis.wright.edu/library/download/SPOT-Provenance_Aware_LSD.pdf Sensor] | ||
| + | **[[Health Care]] | ||
| + | * [http://knoesis1.wright.edu/library/ontologies/provenir/provenir.owl Download Provenir ontology schema (OWL file)]* | ||
| + | }} | ||
| + | <!--end section right 1 --> | ||
| + | |||
| + | <!--start section right 2 --> | ||
| + | |- | ||
| + | | | ||
| + | {{block | ||
| + | |title= <span style="color:#ffffff">Citing Provenir ontology</span> | ||
| + | |title_background=#556b2f | ||
| + | |content= | ||
| + | Please use the following reference when citing Provenir ontology | ||
| + | * Satya S. Sahoo, Amit Sheth, 'Provenir ontology: Towards a Framework for eScience Provenance Management', Microsoft eScience Workshop, Pittsburgh, PA Oct 15-17, 2009 | ||
| + | }} | ||
| + | <!--end section right 1 --> | ||
==References== | ==References== | ||
1. Sahoo SS, Thomas, C., Sheth, A., York, W. S., and Tartir, S. Knowledge modeling and its application in life sciences: a tale of two ontologies. In: Proceedings of the 15th international Conference on World Wide Web WWW '06 2006 May 23 - 26; Edinburgh, Scotland; 2006. p. 317-326. <br/> | 1. Sahoo SS, Thomas, C., Sheth, A., York, W. S., and Tartir, S. Knowledge modeling and its application in life sciences: a tale of two ontologies. In: Proceedings of the 15th international Conference on World Wide Web WWW '06 2006 May 23 - 26; Edinburgh, Scotland; 2006. p. 317-326. <br/> | ||
2. Smith B, Ceusters W, Klagges B, Kohler J, Kumar A, Lomax J, et al. Relations in biomedical ontologies. Genome Biol 2005;6(5):R46. | 2. Smith B, Ceusters W, Klagges B, Kohler J, Kumar A, Lomax J, et al. Relations in biomedical ontologies. Genome Biol 2005;6(5):R46. | ||
Latest revision as of 00:50, 14 May 2011
|
|