Difference between revisions of "Provenir Ontology"
| Line 1: | Line 1: | ||
| − | <!-- | + | <!--begin table --> |
| + | {| style="width:100%; vertical-align:top;" | ||
| + | <!--begin left panel --> | ||
| + | | style="width:60%; border:1px solid #cef2e0; background:#cef2e0; vertical-align:top; color:#000;" | | ||
| + | {| style="width:100%; vertical-align:top;" | ||
| + | |||
| + | <!--start section left 1 --> | ||
| + | |- | ||
| + | | | ||
{{block | {{block | ||
|title= What is Provenir Ontology (PO)? | |title= What is Provenir Ontology (PO)? | ||
| Line 9: | Line 17: | ||
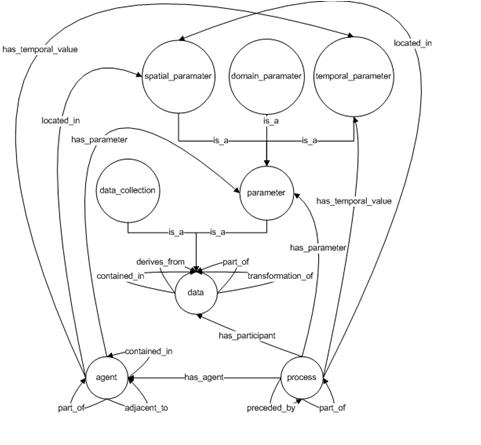
We introduce the provenir ontology as a common provenance model, which forms the core component of a modular approach to provenance management framework in eScience. Domain-specific details are an important component of provenance representation. But, a single monolithic provenance ontology that models all possible details from different domains (biology, marine sciences, and astronomy) is clearly not feasible. Hence, our proposed modular approach involves integrated use of multiple ontologies, each modeling provenance metadata specific to a particular domain (for example, the ProPreO ontology represents proteomics domain-specific provenance[1]). These multiple ontologies will use the provenir ontology as the common reference model, hence making it easier for their associated instances to be interoperable. This modular framework represents a scalable and flexible approach to provenance modeling that can be adapted to the specific requirement of different domains. In the next two Sections, we describe the classes and the properties in the provenir ontology (Figure 1). | We introduce the provenir ontology as a common provenance model, which forms the core component of a modular approach to provenance management framework in eScience. Domain-specific details are an important component of provenance representation. But, a single monolithic provenance ontology that models all possible details from different domains (biology, marine sciences, and astronomy) is clearly not feasible. Hence, our proposed modular approach involves integrated use of multiple ontologies, each modeling provenance metadata specific to a particular domain (for example, the ProPreO ontology represents proteomics domain-specific provenance[1]). These multiple ontologies will use the provenir ontology as the common reference model, hence making it easier for their associated instances to be interoperable. This modular framework represents a scalable and flexible approach to provenance modeling that can be adapted to the specific requirement of different domains. In the next two Sections, we describe the classes and the properties in the provenir ontology (Figure 1). | ||
}} | }} | ||
| − | <!--end section 1 | + | <!-- end section left 1 --> |
| − | + | ||
| − | + | ||
| + | <!--start section left 2 --> | ||
| + | |- | ||
| + | | | ||
{{block | {{block | ||
|title= Classes of Provenir Ontology (PO) | |title= Classes of Provenir Ontology (PO) | ||
| Line 29: | Line 38: | ||
:::* domain_parameter: The domain_parameter class is used to model domain-specific parameters (for example, tolerable salinity levels for ocean buoys). | :::* domain_parameter: The domain_parameter class is used to model domain-specific parameters (for example, tolerable salinity levels for ocean buoys). | ||
}} | }} | ||
| − | <!--end section 2 --> | + | <!-- end section left 2 --> |
| − | <!--start section 3 --> | + | <!--start section left 3 --> |
| − | + | |- | |
| + | | | ||
{{block | {{block | ||
|title= Properties of Provenir Ontology (PO) | |title= Properties of Provenir Ontology (PO) | ||
| Line 50: | Line 60: | ||
:::* located_in – An instance of data or agent is associated with exactly one spatial region that is its exact location at given instance of time. In provenir ontology, this relation has two domain class agent and data_collection with spatial_parameter as range class. | :::* located_in – An instance of data or agent is associated with exactly one spatial region that is its exact location at given instance of time. In provenir ontology, this relation has two domain class agent and data_collection with spatial_parameter as range class. | ||
}} | }} | ||
| − | <!--end section 3 --> | + | <!-- end section left 3 --> |
| + | |||
| + | |}<!-- end left panel --> | ||
| + | |||
| + | <!-- begin right panel --> | ||
| + | | style="width:40%; border:1px solid #ffffcc; background:#f5faff; vertical-align:top;" | <!--right panel --> | ||
| + | {| style="width:100%; vertical-align:top;" | ||
| + | |||
| + | <!--start section right 1 --> | ||
| + | |- | ||
| + | | | ||
| + | {{block | ||
| + | |title= Quick Links | ||
| + | |title_background=#ffffcc | ||
| + | |content= | ||
| + | * [[Applications]] - Applications of Provenir Ontology in different domains | ||
| + | * [[Parasite Life Cycle ontology]] - Ontology that models all the lifecycle stages of various kinetoplastids | ||
| + | * [[Provenir Ontology]] - Upper level provenance ontology | ||
| + | * [[Publications]] - conference and journal publications generated from this research | ||
| + | * [[TcruziSPSE_Team|People]] - people who are involved in this research | ||
| + | }} | ||
| + | <!--end section right 1 --> | ||
| + | |||
| + | |||
==Download== | ==Download== | ||
Revision as of 15:56, 11 June 2010
|
|