Difference between revisions of "Biomedical Sciences"
| Line 31: | Line 31: | ||
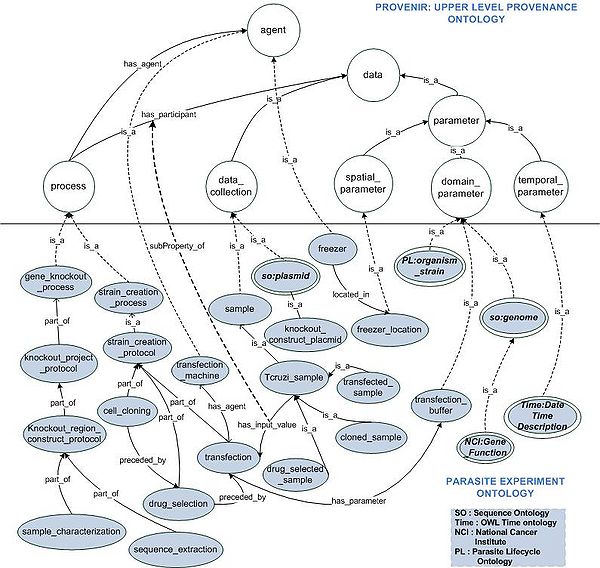
# Modeling of Experimental Processes and Protocols: Two classes namely, gene_knockout_process and strain_creation_process, are created as subclass of provenir:process3 class, to model the generic gene knockout and strain creation processes. The knockout_project_protocol and strain_creation_protocol classes represent the particular protocols used in the lab. The GKO and SP protocols consist of multiple sub-processes, which are also modeled in PEO, for example sequence_extraction, plasmid_construction, transfection, drug_selection, and cell_cloning (Figure 1). | # Modeling of Experimental Processes and Protocols: Two classes namely, gene_knockout_process and strain_creation_process, are created as subclass of provenir:process3 class, to model the generic gene knockout and strain creation processes. The knockout_project_protocol and strain_creation_protocol classes represent the particular protocols used in the lab. The GKO and SP protocols consist of multiple sub-processes, which are also modeled in PEO, for example sequence_extraction, plasmid_construction, transfection, drug_selection, and cell_cloning (Figure 1). | ||
| − | # Modeling of Datasets and Parameters Used in Protocols: A novel feature of the provenir ontology is the distinct modeling of provenir:data_collection (representing entities that undergo processing in an experiment) and provenir:parameter (representing entities that influence the behavior of a process or agent). In the PEO, for example process, transfection, the input value Tcruzi_sample is modeled as a subclass of provenir:data_collection class and the parameter value transfection_buffer is modeled as a sub-class of the provenir:parameter class. Further, the parameter values are also categorized along the space, time, and theme (domain-specific) dimensions, for example the date on which an experiment is conducted is modeled using the Time:DateTimeDescription class (Figure 1). [[Image:ExtendingProvenir.jpg| | + | # Modeling of Datasets and Parameters Used in Protocols: A novel feature of the provenir ontology is the distinct modeling of provenir:data_collection (representing entities that undergo processing in an experiment) and provenir:parameter (representing entities that influence the behavior of a process or agent). In the PEO, for example process, transfection, the input value Tcruzi_sample is modeled as a subclass of provenir:data_collection class and the parameter value transfection_buffer is modeled as a sub-class of the provenir:parameter class. Further, the parameter values are also categorized along the space, time, and theme (domain-specific) dimensions, for example the date on which an experiment is conducted is modeled using the Time:DateTimeDescription class (Figure 1). [[Image:ExtendingProvenir.jpg| center| 600px| Figure: 1 Extension of PO to PEO]] |
| − | # Modeling of Experiment Materials and Other Details: PEO uses provenir:agent class to model researchers, instruments, and biological agents involved in an experiment. For example, transfection_machine, microarray_plate_reader are instruments modeled as subclass of provenir:agent; researcher is an example of human agent; and knockout_plasmid is an example of a biological agent. | + | # Modeling of Experiment Materials and Other Details: PEO uses provenir:agent class to model researchers, instruments, and biological agents involved in an experiment. For example, transfection_machine, microarray_plate_reader are instruments modeled as subclass of provenir:agent; researcher is an example of human agent; and knockout_plasmid is an example of a biological agent. |
In addition to the eleven relationships defined in provenir ontology, new object and datatype properties specific to experiment protocols have been created. For example, four new object properties are defined to model the similarity relationships between two genomic regions, namely is_paralogous_to, is_orthologous_to, is_homologous_to, and is_identical_to. | In addition to the eleven relationships defined in provenir ontology, new object and datatype properties specific to experiment protocols have been created. For example, four new object properties are defined to model the similarity relationships between two genomic regions, namely is_paralogous_to, is_orthologous_to, is_homologous_to, and is_identical_to. | ||
Latest revision as of 18:56, 1 September 2010
|